K. Peter Pauls

Email:
Phone:
Education:
B.Sc. University of Waterloo;
M.Sc. University of Waterloo;
Ph.D. University of Waterloo
Location:
Room:
Our laboratory is working in various areas of plant cell biology, molecular biology and genomics. It has focused on understanding disease-related and developmental phenomena that are of agricultural interest and developing methods to improve crops through biotechnology. Significant research accomplishments of the lab include:
- Sequencing of the genome for bean (Phaseolus vulgaris) variety OAC Rex
- Identification of molecular markers for: bacterial blight resistance in beans, lipoxygenase nulls and low linolenic acid content in soybean, Fusarium graminearum resistance in corn and yield-related traits in beans.
- Development of molecular linkage maps for several bean, soybean and corn breeding populations.
- Development of genetic distance and QTL- index based approaches to marker assisted selection.
- Production of transgenic corn and tomatoes with improved resistance to fungal diseases.
- Development of a genotype-independent Agrobacterium-based transformation system for corn.
- Cloning and sequencing of genes from Brassica microspores that may be involved in the commitment of cells to embryogenesis.
- Cloning and sequencing genes coding for structural and regulatory genes in the phenylpropanoid and folic acid synthesis pathways in Phaseolus.
- Release of new white bean and kidney bean varieties developed through conventional breeding.
- Measurement of the persistence of DNA from transgenic plants in agricultural soils.
News Item:
U of G Research Beans Donated to Local Organizations
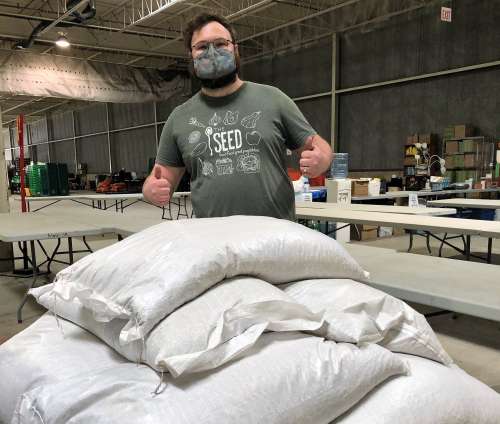 Researchers with the U of G dry bean breeding program recently donated more than 2,000 kilograms of beans to the Guelph Food Bank, the non-profit group The Seed and the University’s Hospitality Services.
Researchers with the U of G dry bean breeding program recently donated more than 2,000 kilograms of beans to the Guelph Food Bank, the non-profit group The Seed and the University’s Hospitality Services.
The beans grew out of a research project led by Dr. Peter Pauls. The four-year study investigated the feasibility and potential benefits of growing a mixture of bean varieties in a single field rather than a single crop. The researchers wanted to know whether the approach would enhance genetic diversity, crop yield and resilience of the resulting crops.
The study, supported by U of G’s Food from Thought program, involved planting and harvesting thousands of bean plants in Woodstock, Ont., and at the Elora research station, said Pauls.
For the full story visit here: https://news.uoguelph.ca/2021/05/u-of-g-research-beans-donated-to-local-...
Courses:
Relevant Links:
Selected Publications:
Reinprecht Y, Schram L, Marsolais F, Smith TH, Hill B and Pauls KP (2020) Effects of Nitrogen Application on Nitrogen Fixation in Common Bean Production. Front. Plant Sci. 11:1172.
Islam, Nishat; Bett, Kirstin; Pauls, K. Peter; Marsolais, Frédéric ; Dhaubhadel, Sangeeta. (2020) Postharvest seed coat darkening in pinto bean (Phaseolus vulgaris) is regulated by Psd, an allele of the basic helix-loop-helix transcription factor P. Plants, People, Planet Plants People Planet. 2020;00:1–15., https://doi.org/10.1002/ppp3.10132
Jiang Y, MacLean DE, Perry GE, Marsolais F, Hill B, Pauls KP. (2020) Evaluation of beneficial and inhibitory effects of nitrate on nodulation and nitrogen fixation in common bean (Phaseolus vulgaris). Legume Science. 2020;e45. https://onlinelibrary.wiley.com/doi/full/10.1002/leg3.45
Erfatpour, M., Pauls, K.P. (2020) A R2R3-MYB gene-based marker for the non-darkening seed coat trait in pinto and cranberry beans (Phaseolus vulgaris L.) derived from ‘Wit-rood boontje’. Theor Appl Genet 133:1977–1994. https://link.springer.com/article/10.1007/s00122-020-03571-7
Wilker, J.W., Navabi, A., Rajcan, I., Marsolais, F., Hill, B., Torkamaneh, D., Pauls, K.P. (2019) Agronomic Performance and Nitrogen Fixation of Heirloom and Conventional Dry Bean Varieties Under Low-Nitrogen Field Conditions. Front. Plant Sci. 10:952. doi: 10.3389/fpls.2019.00952
Monk, JM, Wu, W, Lepp, D, Wellings, HR, Hutchinson, AL, Liddle, DM, Graf, D, Pauls, KP, Robinson, LE, Power KA (2019) Navy bean supplemented high fat diet improves intestinal health, epithelial barrier integrity and critical aspects of the obese inflammatory phenotype. The Journal of Nutritional Biochemistry. 70: 91-104. https://www.sciencedirect.com/science/article/abs/pii/S0955286318311835?via%3Dihub
Corzo, M., Quiñones, M.L, and Pauls. K.P. (2019) First report of Xanthomonas campestris causing black rot of chard in Cuba. New Disease Reports 39: 13. https://www.ndrs.org.uk/article.php?id=039013
Chen, Y., Zhang, H., Liub, R., Mats, L., Zhub, H., Pauls, K.P., Denga, Z., Tsao, R. (2019) Antioxidant and anti-inflammatory polyphenols and peptides of common bean (Phaseolus vulgaris L.) milk and yogurt in Caco-2 and HT-29 cell models. J. Functional Foods 53: 125–135. https://www.sciencedirect.com/science/article/pii/S175646461830642X?casa_token=t_zeXdj6PaQAAAAA:qwijyvcAe12aDl-cDCme4O4yDcBO8d1q2cDjKNk-03jfnXk4auuvkCD1ifqn4UfQYJI8D9U_lQ
Christoper Dumigan, Gregory Perry, K. Peter Pauls, and Manish Raizada. (2018) Draft Genome Sequence of Enterobacter cloacae 3D9 (Phylum Proteobacteria). Microbiology Resource Announcements https://mra.asm.org/content/7/16/e00902-18
Dumigan, C., Perry, G., Pauls, K.P. and Raizada, M. (2018) Draft Genome Sequence of Enterobacter cloacae. 3F11 (Phylum Proteobacteria). Microbiology Resource Announcements. 7:e00846-18. https://doi.org/10.1128/MRA.00846-18.
Balasubramanian, P.M., Rupert, T., Marsolais, F., Park, S.J., Navabi, A., Smith, T.H., Khanal, R., Burt, A.J., Pauls, K.P. (2018) AAC Argosy navy dry bean. Canadian Journal of Plant Science, 98: 199-202 dx.doi.org/10.1139/cjps-2017-0176
Balasubramanian, P.M., Rupert, T., Marsolais, F., Park, S.J., Navabi, A., Smith, T.H., Khanal, R., Burt, A.J., Pauls, K.P. (2018) AAC Shock navy dry bean. Canadian Journal of Plant Science, 98: 203-206 dx.doi.org/10.1139/cjps-2017-0175
Erfatpour, M., Navabi, A. and Pauls, K.P. (2018) Mapping the non‑darkening trait from ‘Wit‑rood boontje’ in bean (Phaseolus vulgaris). Theor App Gen (2018) 131:1331–1343. DOI:10.1007/s00122-018-3081-y
Farid, M., Earl, H.J., Pauls, K.P. and Navabi, A. (2017) Response to selection for improved nitrogen fixation in common bean (Phaseolus vulgaris L.). Euph 213: 99. doi:10.1007/s10681-017-1885-5
Xie, W., Khanal, R., McClymont, S., Stonehouse, R., Bett, K., Yu, K.F., Pauls, K.P. and Navabi, A. (2017) Interaction of quantitative trait loci for resistance to common bacterial blight and pathogen isolates in Phaseolus vulgaris L. Mol Breed 37:55. doi:10.1007/s11032-017-0657-1
Xie, W., Perry, G., Martin, C., Shim, Y., Navabi, A. and Pauls. K.P. (2017) Molecular Characterization of dihydroneopterin aldolase and aminodeoxychorismate synthase in common bean - genes coding for enzymes in the folate synthesis pathway. Genome 60: 588–600. DOI:10.1139/gen-2016-0227
Freixas-Coutin, J.A., Munholland, S., Silva, A., Subedi, S., Crosby, W.L., Pauls, K.P. and Bozzo, G.G. (2017) Proanthocyanidin accumulation and transcriptional responses in the seed coat of cranberry beans (Phaseolus vulgaris L.) with different susceptibility to postharvest darkening. BMC Plant Biol 17:89. DOI 10.1186/s12870-017-1037-z
Ahad, R., Zhou, T., Lepp, D. and Pauls, K.P. (2017) Draft genome sequence of Citrobacter freundii Strain A47, resistant to the mycotoxin deoxynivalenol. Genome Announc 5:e00019-17. https://doi.org/10.1128/genomeA.00019-17
Ahad, R., Zhou, T., Lepp, D. and Pauls, K.P. (2017) Microbial detoxification of eleven food and feed contaminating trichothecene mycotoxins. BMC Biotech 17:30. DOI 10.1186/s12896-017-0352-7
Chen, P.X., Zhang, H., Marcone, M.F. Pauls, K.P., Liu, R., Tang, Y., Zhang, B., Renaud, J.B. and Tsao, R. (2017) Anti-inflammatory effects of phenolic-rich cranberry bean (Phaseolus vulgaris L.) extracts and enhanced cellular antioxidant enzyme activities in Caco-2 cells. J Functional Foods 38: 675-685. https://doi.org/10.1016/j.jff.2016.12.027
Khanal, R., Smith, T.H., Michaels, T.E. and Pauls, K.P. (2017) Lighthouse Common Bean. Can J Plant Sci. 97: 165-168. 10.1139/cjps-2016-0073
Reinprecht, Y. and Pauls, K.P. (2016) Microsomal omega-3 fatty acid desaturase genes in low linolenic acid soybean line RG10 and validation of major linolenic acid QTL. Front. Genet. 7:38. doi: 10.3389/fgene.2016.00038
Monk, J.M., Lepp, D., Zhang, C.P., Wu, W., Zarepoor, L., Lu, J.T., Tsao, R., Pauls, K.P., Wood, G.E., Robinson, L.E. and Power, K.A. (2015) Diets enriched with cranberry beans alter the microbiota and mitigate colitis severity and associated inflammation. J Nutrition Biochem 28: 129-139. DOI:10.1016/j.jnutbio.2015.10.014
Reinprecht, Y., Arif, M., Simon, L.C. and Pauls, K.P. (2015) Genome regions associated with functional performance of soybean stem fibers in polypropylene thermoplastic composites. PLoS ONE 10(7): e0130371. doi:10.1371/journal.pone.0130371
Zou, X., Shi, C., Austin, R.S., Merico, D., Munholland, S., Marsolais, F., Navabi, A., Crosby, W., Pauls, K.P., Yu, K.-F., and Cui, Y. (2014) Genome-wide single nucleotide polymorphism and insertion-deletion discovery through next-generation sequencing of reduced representation libraries in common bean. Mol Breed 33: 769-778. DOI 10.1007/s11032-013-9997-7
Reinprecht, Y., Z. Yadegari, G.E. Perry, M. Siddiqua, L.C. Wright, P.E. McClean and K.P. Pauls. (2013). In silico comparison of genomic regions containing genes coding for enzymes and transcription factors for the phenylpropanoid pathway in Phaseolus vulgaris L. and Glycine max L.Merr. Frontiers in Plant Science. 4: 317. DOI: 10.3389/fpls.2013.00317.
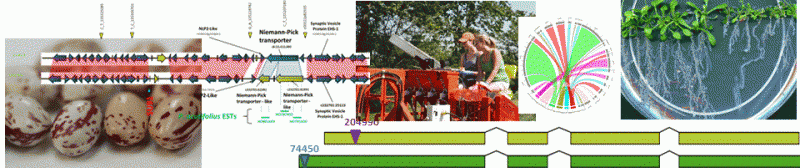
Perry, G.E., C. DiNatale, W. Xie, A. Navabi, Y. Reinprecht, W. Crosby, K. Yu, C. Shi and K.P. Pauls. (2013). A comparison of the molecular organization of genomic regions associated with resistance to common bacterial blight in two Phaseolus vulgaris genotypes. Frontiers in Plant Science 4: 318. DOI: 10.3389/fpls.2013.00318.
Khanal, S., J. Xue, R. Khanal, W. Xie, J. Shi, K.P. Pauls and A. Navabi. (2012). Quantitative Trait Loci Analysis of Folate Content in Dry Beans, Phaseolus vulgaris L. International Journal of Agronomy. DOI: 10.1155/2013/983641.
Smith, T. H., T.E. Michaels, A. Navabi and K.P. Pauls. (2012). OAC Inferno common bean. Canadian Journal Plant Science. 92: 589-592.
Taheri, A., J. Subramanian, J.A. Cline, M.N. Raizada and K.P Pauls. (2012). A WD-repeat gene from peach (Prunus persica L.) is a functional ortholog of Arabidopsis thaliana TRANSPARENT TESTA GLABRA1. In Vitro Cellular & Developmental Biology - Plant. 48: 23–29.
Smith, T. H., T.E. Michaels, A. Navabi and K.P. Pauls. (2012). OAC Inferno common bean. Canadian Journal Plant Science. 92: 589-592.
Xie, W., K.F. Yu, K.P. Pauls and A. Navabi. (2012) Applications of Image Analysis in Studies of Quantitative Disease Resistance, Exemplified Using Common Bacterial Blight – Common Bean Pathosystem. Phytopathology. 102: 434-442.